circRNA Analysis
WANG Ziyi
Date: 07 3月 2023
ENSMUSG00000029661:+:6:4529067:4531010
1. Sequences
| id | gene | transcript | strand | chrom | startGR | endGR | length | seq | type |
|---|---|---|---|---|---|---|---|---|---|
| ENSMUSG00000029661:+:6:4529067:4531010 | ENSMUSG00000029661 | ENSMUST00000031668 | + | 6 | 4529068 | 4531010 | 162 | GGUGAAACUGGUCUCCGAGGUGACACUGGCAACACUGGUAGAGAUGGUGCUCGUGGCAUUCCUGGUGCUGUAGGUGCCCCUGGUCCUGCUGGGGCCUCAGGUGACCGGGGUGAAGCUGGUGCUGCCGGUCCUUCUGGCCCAGCUGGUCCUCGGGGUAGCCCUGGUGAAACUGGUCUCCGAGGUGACACUGGCAACACUGGUAGAGAUGGUGC | circ |
| ENSMUSG00000029661:+:6:4529067:4531010 | ENSMUSG00000029661 | ENSMUST00000031668 | + | 6 | 4529068 | 4531010 | 22 | GGGGUAGCCCUGGUGAAACUGG | bsj |
| ENSMUSG00000029661:+:6:4529067:4531010 | ENSMUSG00000029661 | ENSMUST00000031668 | + | 6 | 4528868 | 4529077 | 210 | CUCUGAAAGGGGCCAGAAAAGAUGAACUUCCGACUGUUUUUCUCAUUCAACAUCCUACAAAGUGGUGGGAAACUGAAAAACACUGAAAUUCUAAAAGGGUGGCCAGCAUUAUAGAAAUUUUUGUCAAAGCACUUUGCAGUUCAUUUCCACUCAGAAGUUGAGCAGGAAAGCAAAAUGUGGCUCUUGCUUUGUACUUCCAGGGUGAAACUG | ie_up |
| ENSMUSG00000029661:+:6:4529067:4531010 | ENSMUSG00000029661 | ENSMUST00000031668 | + | 6 | 4531001 | 4531210 | 210 | GGGUAGCCCUGUAAGUAGAAACCACAGCUUUUUGGAUAUUCAUAGUUCAUGGUUUACAGACAUCUGAGAGGCUGUGAGUGUGAUGUCUUUAUCUCCCACCUAGGGUGAACGUGGUGAAGUUGGCCCUGCUGGCCCCAAUGGAUUUGCUGGUCCUGCUGUGAGUAUCAUAUUAAGGGAGAUUAAGAUGACAUCACAGUGAGUCAGAAGGCC | ie_down |
- Note:
- id: unique identifier.
- gene: represents the gene from which the circRNA arises.
- transcript: transcript whose exon coordinates overlap with the detected back-spliced junction coordinate and used in the downstream analysis.
- strand: is the strand from which the gene is transcribed.
- chrom: is the chromosome from which the circRNA is derived.
- startGR: is the 5’ coordinate of the genomic range from which the sequence are extracted.
- endGR: is the 3’ coordinate of the genomic range from which the sequence are extracted.
- length: length of the extracted sequences.
- seq: sequence used in the downstream analysis.
- type: type of sequences retrieved. If type = “circ” the sequences derive from the internal circRNA sequences. If type = “bsj” the sequences derive from the back spliced junctions (BSJ). If type = “ie_up” the Intron or Exon sequences derive from the up-stream of BSJ up to 210 bp. If type = “ie_down” the Intron or Exon sequences derive from the down-stream of BSJ up to 210 bp.
- Please Click HERE to learn more details about the results from circRNAprofiler.
2. RNA Binding Proteins Analysis
RBP on full sequence
Plot
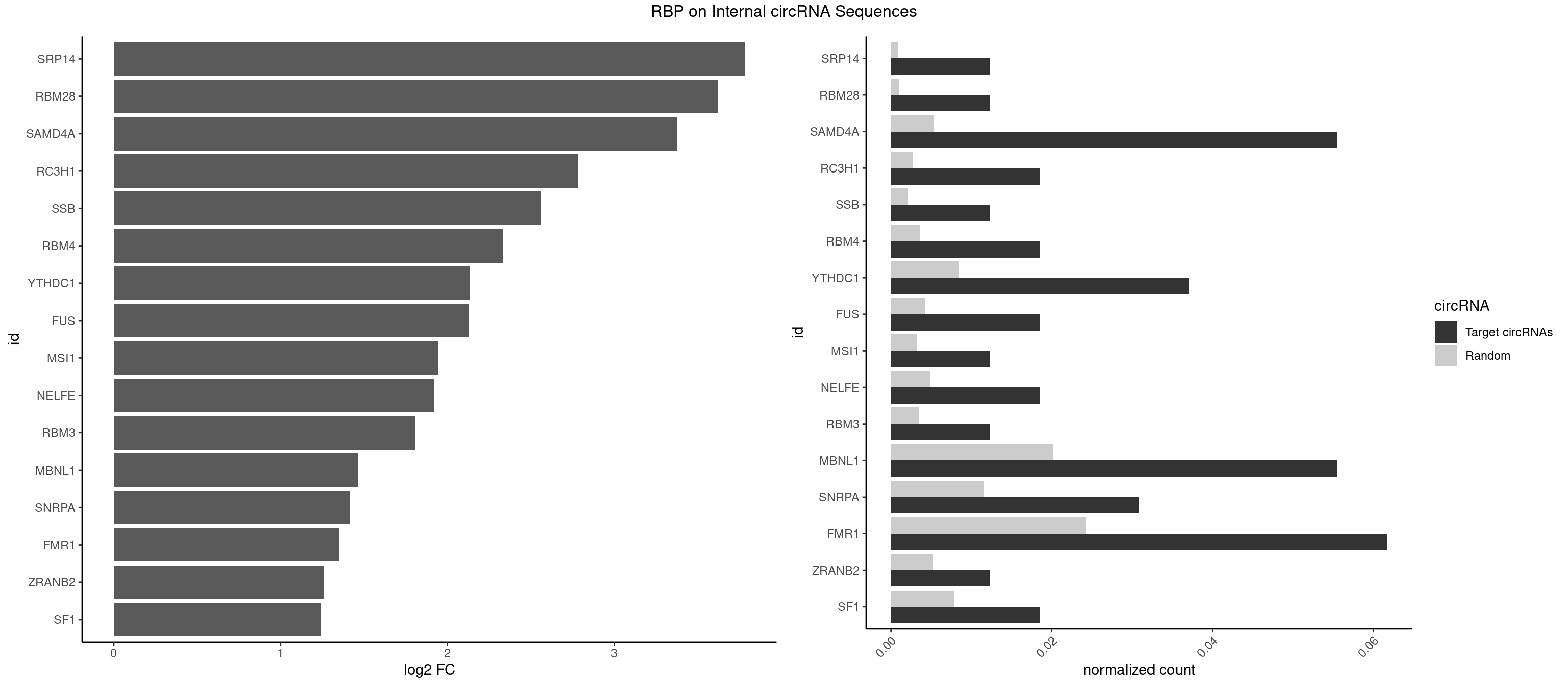
Spreadsheet
| id | foreground | background | foregroundNorm | backgroundNorm | log2FC | motifF | motifB |
|---|---|---|---|---|---|---|---|
| SRP14 | 1 | 32 | 0.01234568 | 0.0008963738 | 3.783762 | CUGUAG | CCUGUA,CGCCUG,CUGUAG,GCCUGU |
| RBM28 | 1 | 36 | 0.01234568 | 0.0010050251 | 3.618703 | UGUAGG | AGUAGA,AGUAGG,AGUAGU,GAGUAG,GUGUAG,UGUAGA,UGUAGG,UGUAGU |
| SAMD4A | 8 | 196 | 0.05555556 | 0.0053510797 | 3.376029 | CUGGCA,CUGGCC,CUGGUA,CUGGUC,GCUGGU | CGGGAA,CGGGAC,CGGGCA,CGGGCC,CGGGUA,CGGGUC,CUGGAA,CUGGAC,CUGGCA,CUGGCC,CUGGUA,CUGGUC,GCGGGA,GCGGGC,GCGGGU,GCUGGA,GCUGGC,GCUGGU |
| RC3H1 | 2 | 98 | 0.01851852 | 0.0026891213 | 2.783762 | CCUUCU,CUUCUG | CCCUUC,CCUUCU,CUUCUG,UCCCUU,UCUGUG,UUCUGU |
| SSB | 1 | 76 | 0.01234568 | 0.0020915388 | 2.561370 | UGCUGU | CUGUUU,GCUGUU,UGCUGU,UGUUUU |
| RBM4 | 2 | 134 | 0.01851852 | 0.0036669836 | 2.336303 | CCUUCU,UCCUUC | CCUCUU,CCUUCC,CCUUCU,CGCGCG,CGCGGG,CGCGGU,CUCUUU,CUUCCU,CUUCUU,GCGCGC,GCGCGG,UCCUUC,UUCCUU,UUCUUG |
| YTHDC1 | 5 | 309 | 0.03703704 | 0.0084204808 | 2.136994 | UCCUGC,UGCUGC,UGGUGC | GAAUAC,GAAUGC,GACUAC,GACUGC,GAGUAC,GAGUGC,GCAUAC,GCAUGC,GCCUAC,GCCUGC,GCGUAC,GCGUGC,GGAUAC,GGAUGC,GGCUAC,GGCUGC,GGGUAC,GGGUGC,UAAUAC,UAAUGC,UACUAC,UACUGC,UAGUAC,UAGUGC,UCAUAC,UCAUGC,UCCUAC,UCCUGC,UCGUAC,UCGUGC,UGAUAC,UGAUGC,UGCUAC,UGCUGC,UGGUAC,UGGUGC |
| FUS | 2 | 155 | 0.01851852 | 0.0042374032 | 2.127716 | GGGUGA,UGGUGA | AAAAAA,CGGUGA,CGGUGG,GGGGGG,GGGUGA,GGGUGC,GGGUGG,GGGUGU,UGGUGA,UGGUGG,UGGUGU,UUUUUU |
| MSI1 | 1 | 117 | 0.01234568 | 0.0032052153 | 1.945513 | UAGGUG | AGGAAG,AGGAGG,AGGUAG,AGGUGG,AGUAAG,AGUAGG,AGUUAG,AGUUGG,UAGGAA,UAGGAG,UAGGUA,UAGGUG,UAGUAA,UAGUUA,UAGUUG |
| NELFE | 2 | 179 | 0.01851852 | 0.0048893114 | 1.921265 | GGUCUC,UCUGGC | CUCUCU,CUCUGG,CUGGCU,CUGGUU,GCUAAC,GGCUAA,GGUCUC,GGUUAG,GUCUCU,UCUCUC,UCUCUG,UCUGGC,UCUGGU,UGGCUA,UGGUUA |
| RBM3 | 1 | 129 | 0.01234568 | 0.0035311694 | 1.805788 | GAAACU | AAAACG,AAAACU,AAACGA,AAACUA,AAGACG,AAGACU,AAUACG,AAUACU,AGACGA,AGACUA,AUACGA,AUACUA,GAAACG,GAAACU,GAGACG,GAGACU,GAUACG,GAUACU |
| MBNL1 | 8 | 740 | 0.05555556 | 0.0201276654 | 1.464751 | CCUGCU,CUGCCG,CUGCUG,GUGCUC,GUGCUG,UGCUGC,UGCUGU | ACGCUC,ACGCUG,ACGCUU,AUGCCA,AUGCCC,AUGCCG,AUGCUC,AUGCUU,CCCGCU,CCGCUA,CCGCUC,CCGCUG,CCGCUU,CCUGCU,CGCUGC,CGCUGU,CGCUUC,CGCUUG,CGCUUU,CUCGCU,CUGCCA,CUGCCC,CUGCCG,CUGCCU,CUGCGG,CUGCUA,CUGCUC,CUGCUG,CUGCUU,CUUGCU,CUUGUG,GCGCUC,GCGCUG,GCGCUU,GCUGCG,GCUGCU,GCUUGC,GCUUGU,GCUUUU,GGCUUU,GUCUCG,GUGCUA,GUGCUC,GUGCUG,GUGCUU,UCCGCU,UCGCUA,UCGCUG,UCGCUU,UCUCGC,UCUGCU,UGCGGC,UGCUGC,UGCUGU,UGCUUC,UGCUUU,UGUCUC,UUCGCU,UUGCCA,UUGCCC,UUGCCG,UUGCCU,UUGCUA,UUGCUC,UUGCUG,UUGCUU,UUGUGC,UUUGCU |
| SNRPA | 4 | 425 | 0.03086420 | 0.0115713704 | 1.415375 | AUUCCU,CCUGCU,UCCUGC,UUCCUG | ACCUGC,AGCAGU,AGGAGA,AGUAGG,AGUAGU,AUACCU,AUGCAC,AUGCUG,AUUCCU,AUUGCA,CAGUAG,CCUGCU,CUGCUA,GAGCAG,GAUACC,GAUUCC,GCAGUA,GGAGAU,GGGUAU,GGUAUG,GUAGGC,GUAGUG,GUAUGC,GUUACC,GUUUCC,UACCUG,UAUGCU,UCCUGC,UGCACA,UGCACC,UGCACG,UUACCU,UUCCUG,UUGCAC,UUUCCU |
| FMR1 | 9 | 891 | 0.06172840 | 0.0242292544 | 1.349184 | ACUGGU,AGCUGG,CAGCUG,CUGGUG,GCUGGG | AAAAAA,AAGCGG,AAGGAA,AAGGAG,AAGGAU,AAGGGA,ACUAAG,ACUGGG,ACUGGU,AGCAGC,AGCGAA,AGCGAC,AGCGGC,AGCUGG,AGGAAG,AGGAAU,AGGAGG,AGGAGU,AGGAUG,AGGAUU,AGGCAC,AGGGAG,AGUGGC,AUAGGC,AUGGAG,CAGCUG,CAUAGG,CGAAGG,CGACUG,CGGCUG,CUAAGG,CUAUGG,CUGAGG,CUGGGC,CUGGUG,GAAGGA,GAAGGG,GACAAG,GACAGG,GACUAA,GACUGG,GAGCGA,GAGCGG,GAGGAU,GCAGCU,GCAUAG,GCGAAG,GCGACU,GCGGCU,GCUAAG,GCUAUG,GCUGAG,GCUGGG,GGACAA,GGACAG,GGACUA,GGCAUA,GGCGAA,GGCUAA,GGCUAU,GGCUGA,GGCUGG,GGGCAU,GGGCGA,GGGCUA,GGGCUG,GGGGGG,GUGCGA,GUGCGG,GUGGCU,UAAGGA,UAGCAG,UAGCGG,UAGGCA,UAGUGG,UAUGGA,UGACAA,UGACAG,UGAGGA,UGCGAC,UGCGGC,UGGAGU,UGGCUG,UUUUUU |
| ZRANB2 | 1 | 189 | 0.01234568 | 0.0051609398 | 1.258300 | GGGUGA | AAAGGU,AGGGAA,AGGUAA,AGGUAC,AGGUAG,AGGUAU,AGGUCU,AGGUUA,AGGUUU,CGGUAG,CGGUAU,CGGUCU,GAGGUU,GGGUAA,GGGUGA,GGUAAA,GGUGGU,GUAAAG,GUGGUG,UAAAGG,UGGUAA,UGGUGG |
| SF1 | 2 | 288 | 0.01851852 | 0.0078500611 | 1.238193 | GCUGCC,UGCUGC | ACAGAC,ACAGUC,ACCGAC,ACGAAC,ACUAAC,ACUAAG,ACUAAU,ACUAGC,ACUGAC,ACUUAU,AGUAAC,AGUAAG,AUACUA,AUUAAC,CACAGA,CACCGA,CACUGA,CAGUCA,CGCUGA,CUAACA,GACUAA,GCUAAC,GCUGAC,GCUGCC,UAACAA,UACGAA,UACUAA,UACUAG,UACUGA,UAGUAA,UAUACU,UGCUAA,UGCUGA,UGCUGC |
RBP on BSJ (Exon Only)
Plot
No results under this category.
Spreadsheet
No results under this category.
RBP on BSJ (Exon and Intron)
Plot
No results under this category.
Spreadsheet
No results under this category.
Help
- Note:
- foreground: number of motifs found in the foreground target sequences (e.g. predicted circRNAs).
- background: number of motifs found in the background target sequences (e.g. randomly generated circRNAs).
- foregroundNorm: number of motifs found in the foreground target sequences (+1) divided by the number (or length) of sequences analyzed.
- backgroundNorm: number of motifs found in background target sequences (+1) divided by the number (or length) of sequences analyzed.
- log2FC: log2 fold change calculated in this way (foregroundNorm)/(backgroundNorm).
- motifF: motifs of the corresponding RBP found in the foreground sequences.
- motifB: motifs of the corresponding RBP found in the background sequences.