Isoforom Switch Analysis
WANG Ziyi
Date: 07 3月 2023
ENSMUSG00000069601
1 Isoform Switch
Summary Plot
Only the isoforms with fraction > 5% are shown below.
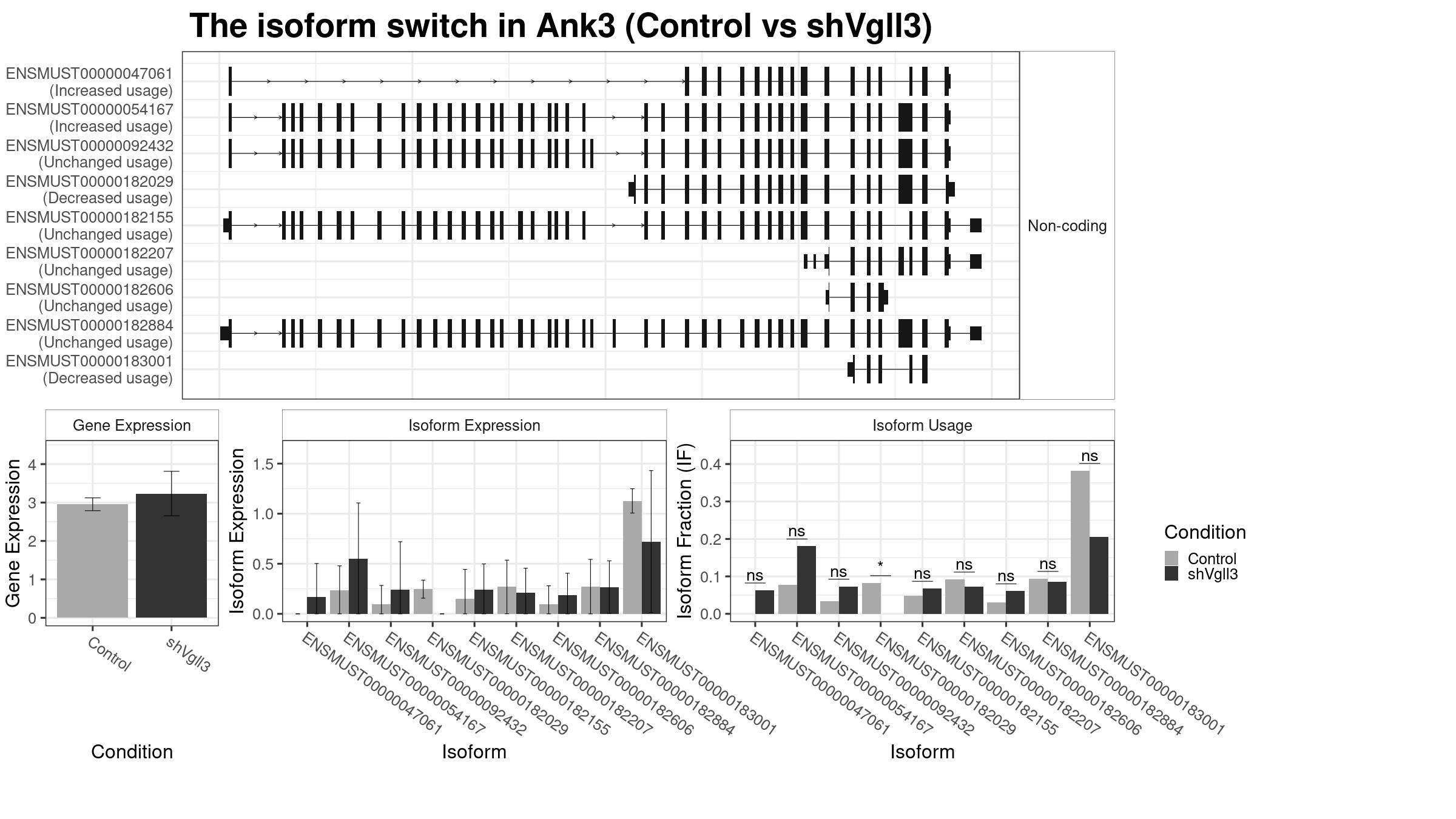
Note:
About the Coding Potential: Please check the spreadsheet in the “Results” tag to confirm the coding potential. The above figure only show the predicted coding potential. The “gene_biotype” in the result spreadsheet is from the database record. Therefore, if you find conflicts between “gene_biotype” and “codingPotential”, please trust the “gene_biotype”.
Notes on IDR and IDR_w_binding: As the prediction of Intrinsically Disordered Regions (IDR) and Intrinsically Disordered Binding Regions (IDBR) are done by two different tools we require two things to annotate the Intrinsically Disordered Binding Regions (IDBR). Firstly the IDBR (predicted by ANCHOR2) must be a region of at least 15 amino acids (after the smoothing). Secondly the fraction of the IDBR which overlaps the IDR predictions (done by IUPred3, again after smoothing) must be at least 80%. When that is the case the IDR type will be annotated as “IDR_w_binding_region” instead of just “IDR”. The current default parameters have not been rigorously tested and should be considered experimental.
For the function of any other domain, please Click HERE to search it through Pfam database.
Results
| isoform_id | gene_id | condition_1 | condition_2 | gene_name | gene_biotype | iso_biotype | gene_overall_mean | gene_value_1 | gene_value_2 | gene_stderr_1 | gene_stderr_2 | gene_log2_fold_change | iso_overall_mean | iso_value_1 | iso_value_2 | iso_stderr_1 | iso_stderr_2 | iso_log2_fold_change | IF_overall | IF1 | IF2 | dIF | isoform_switch_q_value | gene_switch_q_value | PTC | codingPotential |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ENSMUST00000047061 | ENSMUSG00000069601 | Control | shVgll3 | Ank3 | protein_coding | protein_coding | 3.09397 | 2.954539 | 3.233401 | 0.08529314 | 0.2955814 | 0.1296994 | 0.08487345 | 0.00000000 | 0.1697469 | 0.00000000 | 0.1697469 | 4.1678951 | 0.03160000 | 0.00000000 | 0.06320000 | 0.063200000 | 0.970665669 | 0.009096573 | FALSE | |
| ENSMUST00000054167 | ENSMUSG00000069601 | Control | shVgll3 | Ank3 | protein_coding | protein_coding | 3.09397 | 2.954539 | 3.233401 | 0.08529314 | 0.2955814 | 0.1296994 | 0.39133005 | 0.23535154 | 0.5473086 | 0.12431698 | 0.2858045 | 1.1836260 | 0.12916667 | 0.07763333 | 0.18070000 | 0.103066667 | 0.970665669 | 0.009096573 | FALSE | |
| ENSMUST00000092432 | ENSMUSG00000069601 | Control | shVgll3 | Ank3 | protein_coding | protein_coding | 3.09397 | 2.954539 | 3.233401 | 0.08529314 | 0.2955814 | 0.1296994 | 0.16950078 | 0.09575432 | 0.2432472 | 0.09575432 | 0.2432472 | 1.2598301 | 0.05388333 | 0.03436667 | 0.07340000 | 0.039033333 | 1.000000000 | 0.009096573 | FALSE | |
| ENSMUST00000182029 | ENSMUSG00000069601 | Control | shVgll3 | Ank3 | protein_coding | protein_coding | 3.09397 | 2.954539 | 3.233401 | 0.08529314 | 0.2955814 | 0.1296994 | 0.12330125 | 0.24660251 | 0.0000000 | 0.04604567 | 0.0000000 | -4.6814634 | 0.04140000 | 0.08280000 | 0.00000000 | -0.082800000 | 0.009096573 | 0.009096573 | FALSE | |
| ENSMUST00000182155 | ENSMUSG00000069601 | Control | shVgll3 | Ank3 | protein_coding | protein_coding | 3.09397 | 2.954539 | 3.233401 | 0.08529314 | 0.2955814 | 0.1296994 | 0.19512329 | 0.14987761 | 0.2403690 | 0.14987761 | 0.1315922 | 0.6470879 | 0.05831667 | 0.04886667 | 0.06776667 | 0.018900000 | 0.970665669 | 0.009096573 | FALSE | |
| ENSMUST00000182207 | ENSMUSG00000069601 | Control | shVgll3 | Ank3 | protein_coding | protein_coding | 3.09397 | 2.954539 | 3.233401 | 0.08529314 | 0.2955814 | 0.1296994 | 0.23819604 | 0.26851819 | 0.2078739 | 0.13661887 | 0.1265807 | -0.3542782 | 0.08270000 | 0.09240000 | 0.07300000 | -0.019400000 | 1.000000000 | 0.009096573 | FALSE | |
| ENSMUST00000182884 | ENSMUSG00000069601 | Control | shVgll3 | Ank3 | protein_coding | protein_coding | 3.09397 | 2.954539 | 3.233401 | 0.08529314 | 0.2955814 | 0.1296994 | 0.26912179 | 0.27131205 | 0.2669315 | 0.13899209 | 0.1338265 | -0.0226420 | 0.08968333 | 0.09420000 | 0.08516667 | -0.009033333 | 1.000000000 | 0.009096573 | FALSE | |
| ENSMUST00000183001 | ENSMUSG00000069601 | Control | shVgll3 | Ank3 | protein_coding | retained_intron | 3.09397 | 2.954539 | 3.233401 | 0.08529314 | 0.2955814 | 0.1296994 | 0.92467329 | 1.12764039 | 0.7217062 | 0.06203407 | 0.3614664 | -0.6367083 | 0.29423333 | 0.38246667 | 0.20600000 | -0.176466667 | 0.970665669 | 0.009096573 | FALSE |
- Note:
- The comparisons made can be identified as “from ‘condition_1’ to ‘condition_2’”, meaning ‘condition_1’ is considered the ground state and ‘condition_2’ the changed state. This also means that a positive dIF value indicates that the isoform usage is increased in ‘condition_2’ compared to ‘condition_1’. Since the ‘isoformFeatures’ entry is the most relevant part of the switchAnalyzeRlist object, the most-used standard methods have also been implemented to work directly on isoformFeatures.
Help
Please Click HERE to learn more details about the results of IsoformSwitchAnalyzeR.
2 Differential Exon Usage
Counts
All exons whithin this gene region are shown and numbering below.
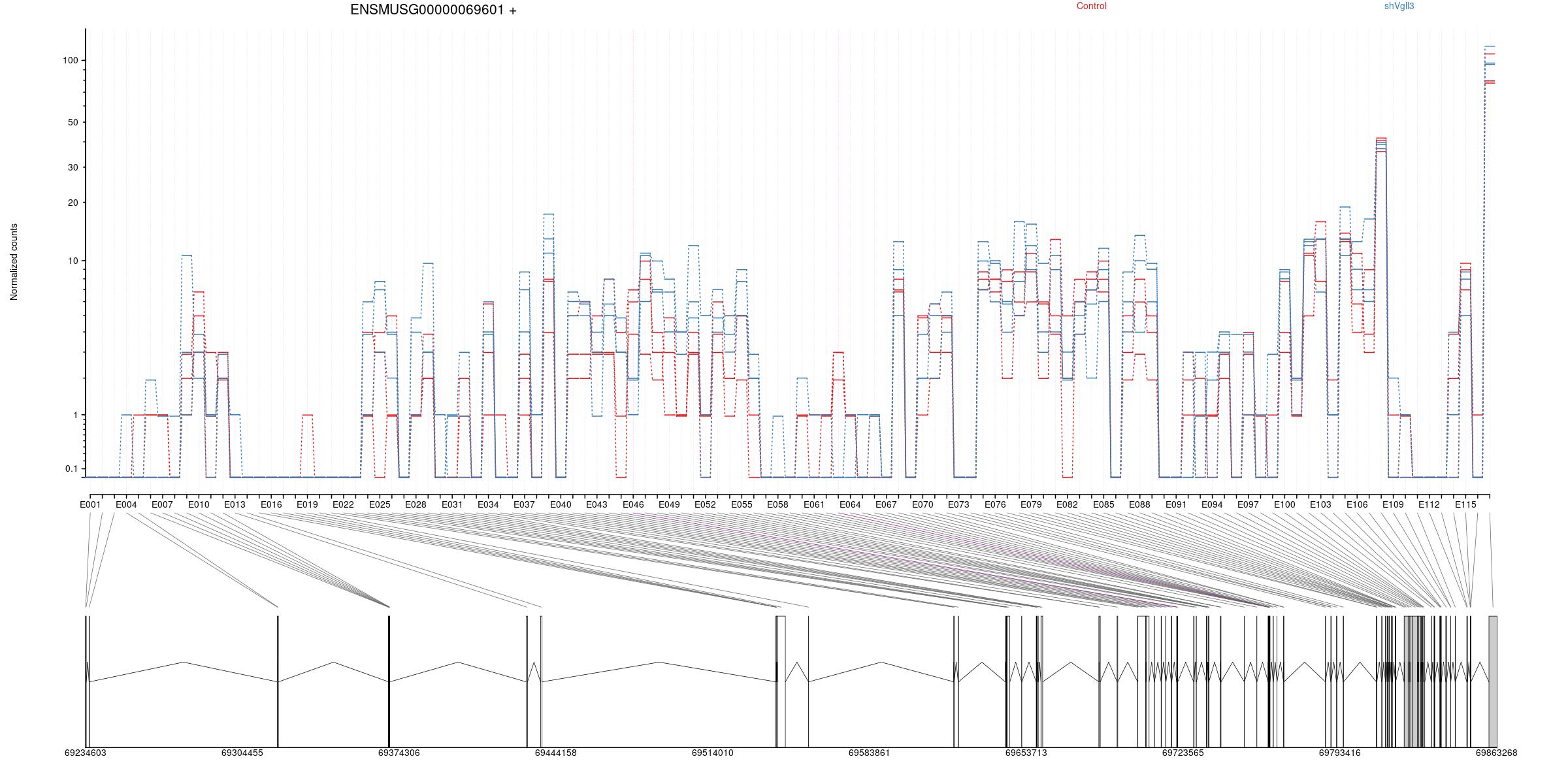
Expression
All exons whithin this gene region are shown and numbering below.
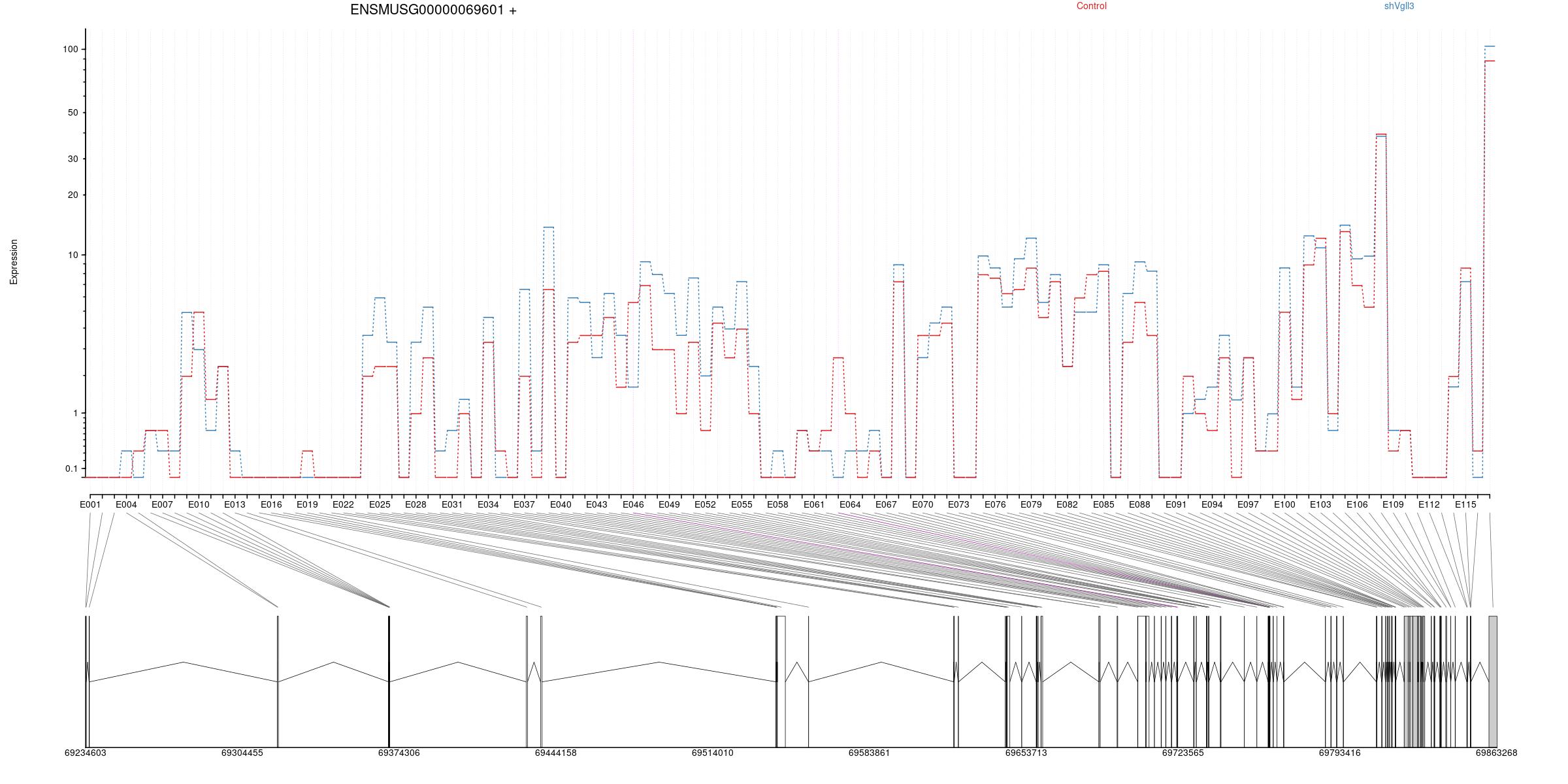
Splicing
All exons whithin this gene region are shown and numbering below.
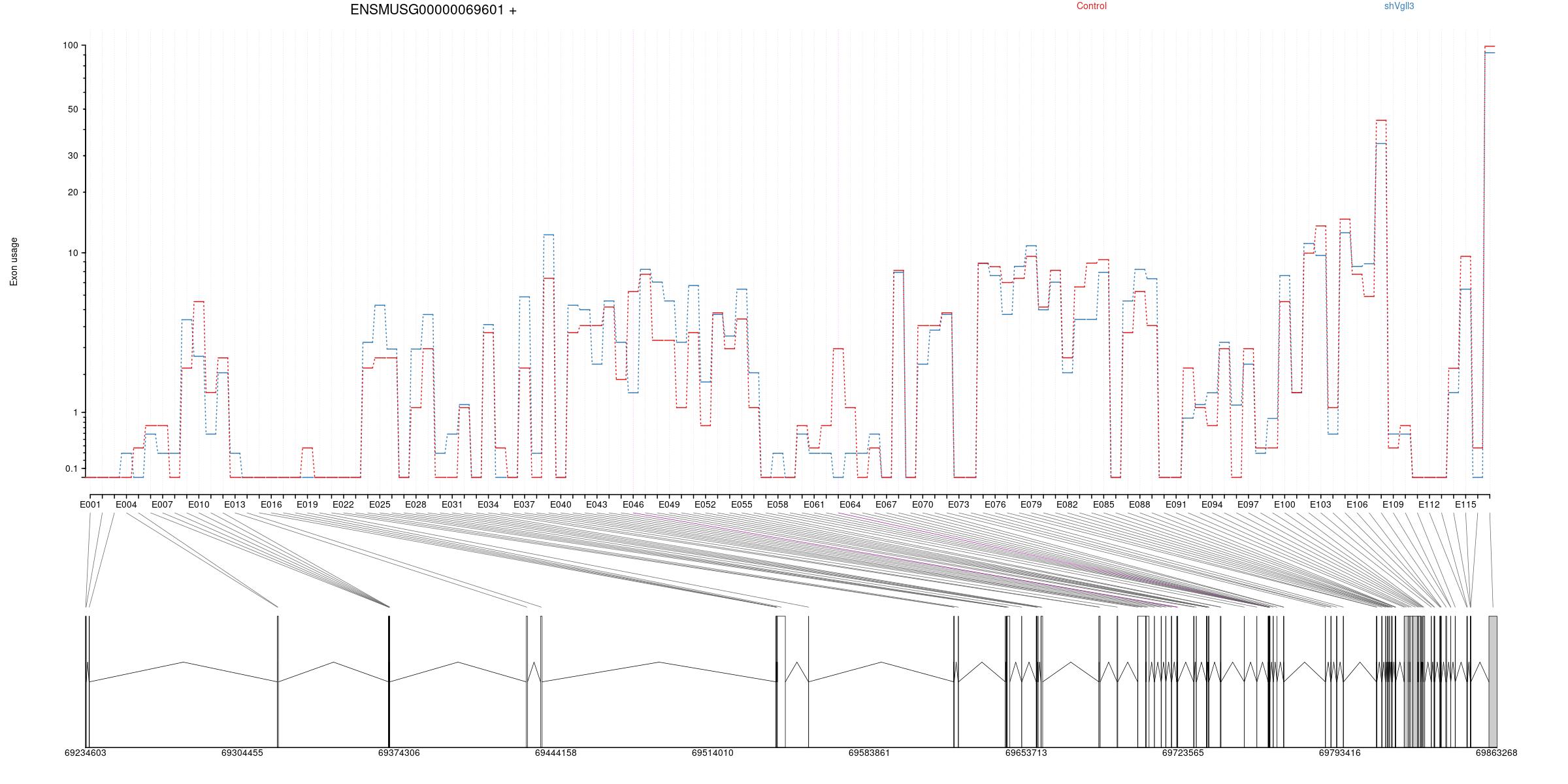
Transcripts
All isoforms whithin this gene region are shown below.
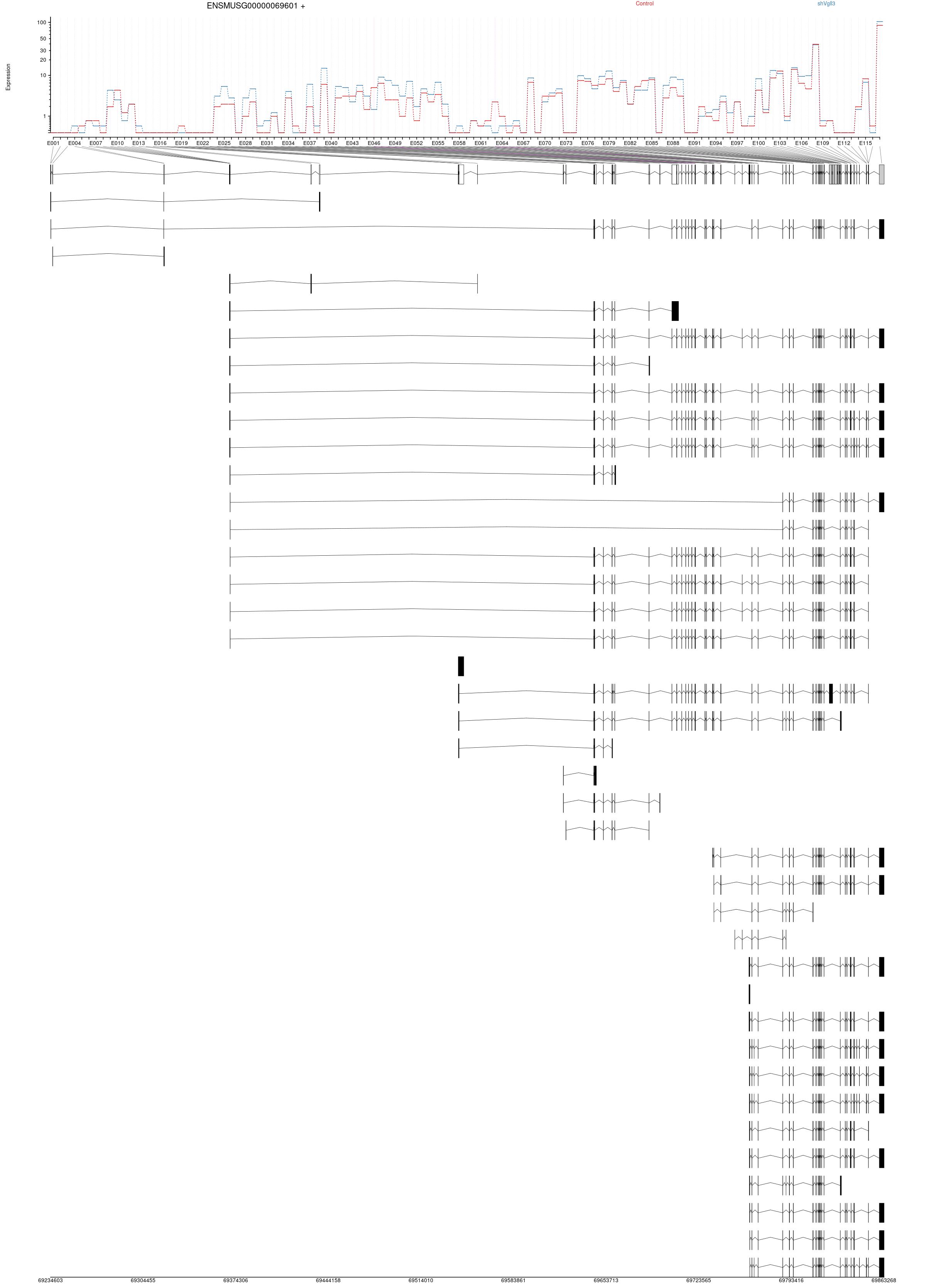

Results
| groupID | featureID | exonBaseMean | dispersion | pvalue | padj | seqnames | start | end | width | strand | shVgll3 | Control | log2fold_Control_shVgll3 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ENSMUSG00000069601 | E001 | 0.0000000 | 10 | 69234603 | 69234825 | 223 | + | ||||||
| ENSMUSG00000069601 | E002 | 0.0000000 | 10 | 69234826 | 69234959 | 134 | + | ||||||
| ENSMUSG00000069601 | E003 | 0.0000000 | 10 | 69236219 | 69236366 | 148 | + | ||||||
| ENSMUSG00000069601 | E004 | 0.1660866 | 0.0348030223 | 0.5947907115 | 10 | 69320059 | 69320097 | 39 | + | 0.112 | 0.000 | -10.722 | |
| ENSMUSG00000069601 | E005 | 0.1657302 | 0.0345837709 | 0.3946360947 | 10 | 69320098 | 69320553 | 456 | + | 0.000 | 0.136 | 13.043 | |
| ENSMUSG00000069601 | E006 | 0.6543404 | 0.0314502366 | 0.8185473652 | 10 | 69369602 | 69369644 | 43 | + | 0.201 | 0.240 | 0.330 | |
| ENSMUSG00000069601 | E007 | 0.4929870 | 0.0218253718 | 0.4424174074 | 10 | 69369645 | 69369672 | 28 | + | 0.112 | 0.240 | 1.330 | |
| ENSMUSG00000069601 | E008 | 0.1613534 | 0.0340373044 | 0.5959013816 | 10 | 69369673 | 69369676 | 4 | + | 0.112 | 0.000 | -13.458 | |
| ENSMUSG00000069601 | E009 | 3.4220720 | 0.1256932920 | 0.3807914510 | 0.88103212 | 10 | 69369677 | 69369894 | 218 | + | 0.731 | 0.507 | -0.987 |
| ENSMUSG00000069601 | E010 | 3.9350196 | 0.0049767255 | 0.0778723485 | 0.59037637 | 10 | 69369895 | 69370037 | 143 | + | 0.561 | 0.815 | 1.066 |
| ENSMUSG00000069601 | E011 | 0.9868397 | 0.0168642319 | 0.2781960138 | 0.82303307 | 10 | 69370038 | 69370051 | 14 | + | 0.201 | 0.394 | 1.330 |
| ENSMUSG00000069601 | E012 | 2.3019951 | 0.0072754271 | 0.6738405959 | 0.96164256 | 10 | 69370052 | 69370114 | 63 | + | 0.485 | 0.554 | 0.330 |
| ENSMUSG00000069601 | E013 | 0.1667494 | 0.0355717341 | 0.5944115627 | 10 | 69430880 | 69431489 | 610 | + | 0.112 | 0.000 | -13.455 | |
| ENSMUSG00000069601 | E014 | 0.0000000 | 10 | 69437283 | 69437968 | 686 | + | ||||||
| ENSMUSG00000069601 | E015 | 0.0000000 | 10 | 69542126 | 69542199 | 74 | + | ||||||
| ENSMUSG00000069601 | E016 | 0.0000000 | 10 | 69542200 | 69542303 | 104 | + | ||||||
| ENSMUSG00000069601 | E017 | 0.0000000 | 10 | 69542304 | 69542307 | 4 | + | ||||||
| ENSMUSG00000069601 | E018 | 0.0000000 | 10 | 69542308 | 69542755 | 448 | + | ||||||
| ENSMUSG00000069601 | E019 | 0.1657302 | 0.0345837709 | 0.3946360947 | 10 | 69542756 | 69546306 | 3551 | + | 0.000 | 0.136 | 13.043 | |
| ENSMUSG00000069601 | E020 | 0.0000000 | 10 | 69556629 | 69556690 | 62 | + | ||||||
| ENSMUSG00000069601 | E021 | 0.0000000 | 10 | 69621344 | 69621354 | 11 | + | ||||||
| ENSMUSG00000069601 | E022 | 0.0000000 | 10 | 69621355 | 69621502 | 148 | + | ||||||
| ENSMUSG00000069601 | E023 | 0.0000000 | 10 | 69623286 | 69623434 | 149 | + | ||||||
| ENSMUSG00000069601 | E024 | 2.7995722 | 0.0595199960 | 0.5161853877 | 0.93149289 | 10 | 69644246 | 69644347 | 102 | + | 0.626 | 0.507 | -0.544 |
| ENSMUSG00000069601 | E025 | 4.1143123 | 0.0095077653 | 0.1040840652 | 0.64156759 | 10 | 69644703 | 69644801 | 99 | + | 0.798 | 0.554 | -1.033 |
| ENSMUSG00000069601 | E026 | 2.7993594 | 0.0172385899 | 0.8004467711 | 0.98165184 | 10 | 69645001 | 69645099 | 99 | + | 0.595 | 0.554 | -0.185 |
| ENSMUSG00000069601 | E027 | 0.0000000 | 10 | 69645100 | 69646295 | 1196 | + | ||||||
| ENSMUSG00000069601 | E028 | 2.1310424 | 0.0085070927 | 0.1165617985 | 0.66203098 | 10 | 69651525 | 69651623 | 99 | + | 0.595 | 0.324 | -1.407 |
| ENSMUSG00000069601 | E029 | 3.9214939 | 0.0312378339 | 0.3469088342 | 0.87431815 | 10 | 69658038 | 69658223 | 186 | + | 0.755 | 0.597 | -0.669 |
| ENSMUSG00000069601 | E030 | 0.1667494 | 0.0355717341 | 0.5944115627 | 10 | 69658224 | 69658465 | 242 | + | 0.112 | 0.000 | -13.455 | |
| ENSMUSG00000069601 | E031 | 0.3274400 | 0.0257975737 | 0.2680516485 | 10 | 69658908 | 69658931 | 24 | + | 0.201 | 0.000 | -14.362 | |
| ENSMUSG00000069601 | E032 | 1.1526198 | 0.0964390073 | 0.9529504085 | 1.00000000 | 10 | 69660148 | 69660246 | 99 | + | 0.337 | 0.324 | -0.086 |
| ENSMUSG00000069601 | E033 | 0.0000000 | 10 | 69660247 | 69660911 | 665 | + | ||||||
| ENSMUSG00000069601 | E034 | 3.9413019 | 0.0172592309 | 0.8011441265 | 0.98165184 | 10 | 69685936 | 69686034 | 99 | + | 0.708 | 0.671 | -0.157 |
| ENSMUSG00000069601 | E035 | 0.1659033 | 0.0345408760 | 0.3947101960 | 10 | 69686035 | 69686557 | 523 | + | 0.000 | 0.136 | 13.043 | |
| ENSMUSG00000069601 | E036 | 0.0000000 | 10 | 69694050 | 69694351 | 302 | + | ||||||
| ENSMUSG00000069601 | E037 | 4.2639596 | 0.0043198938 | 0.0259938673 | 0.36156511 | 10 | 69703215 | 69703313 | 99 | + | 0.837 | 0.507 | -1.407 |
| ENSMUSG00000069601 | E038 | 0.1667494 | 0.0355717341 | 0.5944115627 | 10 | 69703314 | 69706831 | 3518 | + | 0.112 | 0.000 | -13.455 | |
| ENSMUSG00000069601 | E039 | 10.1815729 | 0.0019525028 | 0.0544081121 | 0.51143353 | 10 | 69706832 | 69707029 | 198 | + | 1.125 | 0.923 | -0.741 |
| ENSMUSG00000069601 | E040 | 0.0000000 | 10 | 69707030 | 69708290 | 1261 | + | ||||||
| ENSMUSG00000069601 | E041 | 4.6060188 | 0.0040295132 | 0.3607254772 | 0.87535004 | 10 | 69710620 | 69710718 | 99 | + | 0.798 | 0.671 | -0.518 |
| ENSMUSG00000069601 | E042 | 4.6105688 | 0.0041000712 | 0.5936784301 | 0.95165770 | 10 | 69713660 | 69713758 | 99 | + | 0.777 | 0.704 | -0.299 |
| ENSMUSG00000069601 | E043 | 3.1369477 | 0.0064255636 | 0.2448248342 | 0.80442163 | 10 | 69715723 | 69715821 | 99 | + | 0.525 | 0.704 | 0.789 |
| ENSMUSG00000069601 | E044 | 5.4387504 | 0.0066266699 | 0.8280014912 | 0.98548395 | 10 | 69718191 | 69718388 | 198 | + | 0.818 | 0.790 | -0.111 |
| ENSMUSG00000069601 | E045 | 2.6297420 | 0.0132277161 | 0.2994558382 | 0.84044841 | 10 | 69720577 | 69720675 | 99 | + | 0.626 | 0.454 | -0.808 |
| ENSMUSG00000069601 | E046 | 3.6233459 | 0.0050183830 | 0.0018681630 | 0.07561858 | 10 | 69720969 | 69721067 | 99 | + | 0.392 | 0.862 | 2.095 |
| ENSMUSG00000069601 | E047 | 8.0718499 | 0.0042392947 | 0.8395502357 | 0.98677151 | 10 | 69728152 | 69728349 | 198 | + | 0.965 | 0.942 | -0.085 |
| ENSMUSG00000069601 | E048 | 5.4413085 | 0.0035433896 | 0.0456585286 | 0.47552357 | 10 | 69729275 | 69729373 | 99 | + | 0.906 | 0.635 | -1.086 |
| ENSMUSG00000069601 | E049 | 4.5959890 | 0.0051478556 | 0.1963320295 | 0.77147870 | 10 | 69733871 | 69733960 | 90 | + | 0.818 | 0.635 | -0.749 |
| ENSMUSG00000069601 | E050 | 2.3085838 | 0.0075230709 | 0.0764135908 | 0.58710222 | 10 | 69733961 | 69733969 | 9 | + | 0.626 | 0.324 | -1.545 |
| ENSMUSG00000069601 | E051 | 5.4497218 | 0.0357488030 | 0.1538203067 | 0.72235646 | 10 | 69734058 | 69734156 | 99 | + | 0.890 | 0.671 | -0.873 |
| ENSMUSG00000069601 | E052 | 1.3272831 | 0.3024100896 | 0.4854608376 | 0.92538469 | 10 | 69734824 | 69734840 | 17 | + | 0.442 | 0.240 | -1.263 |
| ENSMUSG00000069601 | E053 | 4.7771546 | 0.0041158659 | 0.9590082818 | 1.00000000 | 10 | 69734841 | 69734898 | 58 | + | 0.756 | 0.763 | 0.030 |
| ENSMUSG00000069601 | E054 | 3.2986180 | 0.0062556043 | 0.6993310711 | 0.96475394 | 10 | 69734899 | 69734919 | 21 | + | 0.655 | 0.597 | -0.255 |
| ENSMUSG00000069601 | E055 | 5.6006135 | 0.0034016469 | 0.2901909124 | 0.83665585 | 10 | 69740028 | 69740100 | 73 | + | 0.873 | 0.735 | -0.545 |
| ENSMUSG00000069601 | E056 | 1.6472690 | 0.0099695703 | 0.3595041553 | 0.87517695 | 10 | 69740278 | 69740340 | 63 | + | 0.485 | 0.324 | -0.893 |
| ENSMUSG00000069601 | E057 | 0.0000000 | 10 | 69750695 | 69750734 | 40 | + | ||||||
| ENSMUSG00000069601 | E058 | 0.1613534 | 0.0340373044 | 0.5959013816 | 10 | 69756268 | 69756321 | 54 | + | 0.112 | 0.000 | -13.458 | |
| ENSMUSG00000069601 | E059 | 0.0000000 | 10 | 69761325 | 69761361 | 37 | + | ||||||
| ENSMUSG00000069601 | E060 | 0.6607752 | 0.1119433158 | 0.8315527805 | 10 | 69761362 | 69761602 | 241 | + | 0.201 | 0.240 | 0.328 | |
| ENSMUSG00000069601 | E061 | 0.3324796 | 0.0289858995 | 0.8697483169 | 10 | 69761603 | 69761634 | 32 | + | 0.112 | 0.136 | 0.329 | |
| ENSMUSG00000069601 | E062 | 0.4940258 | 0.0220188957 | 0.4436635035 | 10 | 69761635 | 69761648 | 14 | + | 0.112 | 0.240 | 1.329 | |
| ENSMUSG00000069601 | E063 | 1.3179932 | 0.0115082444 | 0.0006457175 | 0.03464956 | 10 | 69761649 | 69761783 | 135 | + | 0.000 | 0.597 | 15.668 |
| ENSMUSG00000069601 | E064 | 0.6592664 | 0.0183762368 | 0.2186112938 | 10 | 69761784 | 69761909 | 126 | + | 0.112 | 0.324 | 1.914 | |
| ENSMUSG00000069601 | E065 | 0.1667494 | 0.0355717341 | 0.5944115627 | 10 | 69761910 | 69761962 | 53 | + | 0.112 | 0.000 | -13.455 | |
| ENSMUSG00000069601 | E066 | 0.4896490 | 0.0219633906 | 0.7065279155 | 10 | 69761963 | 69762021 | 59 | + | 0.201 | 0.136 | -0.671 | |
| ENSMUSG00000069601 | E067 | 0.0000000 | 10 | 69762022 | 69762168 | 147 | + | ||||||
| ENSMUSG00000069601 | E068 | 8.0442821 | 0.0023986292 | 0.9324878577 | 0.99930331 | 10 | 69763525 | 69763648 | 124 | + | 0.951 | 0.960 | 0.034 |
| ENSMUSG00000069601 | E069 | 0.0000000 | 10 | 69765277 | 69765288 | 12 | + | ||||||
| ENSMUSG00000069601 | E070 | 3.1140635 | 0.0091091991 | 0.2460226597 | 0.80442163 | 10 | 69768224 | 69768238 | 15 | + | 0.525 | 0.704 | 0.789 |
| ENSMUSG00000069601 | E071 | 3.9324883 | 0.0107320833 | 0.8826841122 | 0.99265898 | 10 | 69768239 | 69768326 | 88 | + | 0.683 | 0.704 | 0.088 |
| ENSMUSG00000069601 | E072 | 4.7616593 | 0.0038788138 | 0.9555062082 | 1.00000000 | 10 | 69786782 | 69786888 | 107 | + | 0.756 | 0.763 | 0.030 |
| ENSMUSG00000069601 | E073 | 0.0000000 | 10 | 69789270 | 69789302 | 33 | + | ||||||
| ENSMUSG00000069601 | E074 | 0.0000000 | 10 | 69789303 | 69789356 | 54 | + | ||||||
| ENSMUSG00000069601 | E075 | 8.8667868 | 0.0021488161 | 0.9832718607 | 1.00000000 | 10 | 69791756 | 69791980 | 225 | + | 0.991 | 0.994 | 0.008 |
| ENSMUSG00000069601 | E076 | 8.0614395 | 0.0024026770 | 0.7159070690 | 0.96912015 | 10 | 69794748 | 69794902 | 155 | + | 0.936 | 0.977 | 0.153 |
| ENSMUSG00000069601 | E077 | 5.7487115 | 0.0253205469 | 0.3028261018 | 0.84356127 | 10 | 69809552 | 69809628 | 77 | + | 0.756 | 0.903 | 0.577 |
| ENSMUSG00000069601 | E078 | 8.0597732 | 0.0474119675 | 0.7457469664 | 0.97314738 | 10 | 69809629 | 69809763 | 135 | + | 0.978 | 0.923 | -0.208 |
| ENSMUSG00000069601 | E079 | 10.3497948 | 0.0019971612 | 0.6292450554 | 0.95595792 | 10 | 69811878 | 69812085 | 208 | + | 1.074 | 1.025 | -0.179 |
| ENSMUSG00000069601 | E080 | 5.0749491 | 0.0273552729 | 0.9067229141 | 0.99563225 | 10 | 69813543 | 69813639 | 97 | + | 0.777 | 0.790 | 0.050 |
| ENSMUSG00000069601 | E081 | 7.5668587 | 0.0467210049 | 0.7319253552 | 0.97196762 | 10 | 69814301 | 69814529 | 229 | + | 0.905 | 0.960 | 0.207 |
| ENSMUSG00000069601 | E082 | 2.3155860 | 0.0620948139 | 0.7409104191 | 0.97290014 | 10 | 69815148 | 69815191 | 44 | + | 0.485 | 0.554 | 0.331 |
| ENSMUSG00000069601 | E083 | 5.4437896 | 0.0035602229 | 0.2465979312 | 0.80442163 | 10 | 69815192 | 69815273 | 82 | + | 0.733 | 0.883 | 0.593 |
| ENSMUSG00000069601 | E084 | 6.4054795 | 0.0030447108 | 0.0328246677 | 0.40805841 | 10 | 69816102 | 69816152 | 51 | + | 0.733 | 0.994 | 1.008 |
| ENSMUSG00000069601 | E085 | 8.5472151 | 0.0022376450 | 0.5883463724 | 0.95124454 | 10 | 69816153 | 69816224 | 72 | + | 0.951 | 1.010 | 0.219 |
| ENSMUSG00000069601 | E086 | 0.0000000 | 10 | 69816654 | 69816680 | 27 | + | ||||||
| ENSMUSG00000069601 | E087 | 4.7654974 | 0.0039182942 | 0.2881077851 | 0.83443547 | 10 | 69817946 | 69817953 | 8 | + | 0.818 | 0.671 | -0.596 |
| ENSMUSG00000069601 | E088 | 7.3970346 | 0.0264351640 | 0.4599387080 | 0.91682762 | 10 | 69817954 | 69817991 | 38 | + | 0.964 | 0.862 | -0.388 |
| ENSMUSG00000069601 | E089 | 5.9261549 | 0.0031765912 | 0.0941520440 | 0.62900916 | 10 | 69817992 | 69818027 | 36 | + | 0.921 | 0.704 | -0.855 |
| ENSMUSG00000069601 | E090 | 0.0000000 | 10 | 69818028 | 69818132 | 105 | + | ||||||
| ENSMUSG00000069601 | E091 | 0.0000000 | 10 | 69821913 | 69821918 | 6 | + | ||||||
| ENSMUSG00000069601 | E092 | 1.4867809 | 0.1619613421 | 0.2571310537 | 0.81097845 | 10 | 69821919 | 69823678 | 1760 | + | 0.275 | 0.507 | 1.327 |
| ENSMUSG00000069601 | E093 | 1.1571500 | 0.0854844374 | 0.9472631690 | 1.00000000 | 10 | 69823679 | 69824462 | 784 | + | 0.337 | 0.324 | -0.085 |
| ENSMUSG00000069601 | E094 | 1.1502314 | 0.0361024590 | 0.4097374254 | 0.89590796 | 10 | 69824463 | 69825764 | 1302 | + | 0.392 | 0.240 | -0.993 |
| ENSMUSG00000069601 | E095 | 3.1244800 | 0.0055848117 | 0.8472680100 | 0.98686918 | 10 | 69825765 | 69827920 | 2156 | + | 0.626 | 0.597 | -0.130 |
| ENSMUSG00000069601 | E096 | 0.6454136 | 0.6487899912 | 0.3039971372 | 10 | 69827921 | 69828305 | 385 | + | 0.335 | 0.000 | -14.617 | |
| ENSMUSG00000069601 | E097 | 2.6252113 | 0.0079264231 | 0.6529103517 | 0.95668705 | 10 | 69828306 | 69829374 | 1069 | + | 0.525 | 0.597 | 0.330 |
| ENSMUSG00000069601 | E098 | 0.3276328 | 0.0281500841 | 0.8682821372 | 10 | 69829375 | 69829592 | 218 | + | 0.112 | 0.136 | 0.329 | |
| ENSMUSG00000069601 | E099 | 0.6497904 | 0.3361743895 | 0.6096038961 | 10 | 69830133 | 69830201 | 69 | + | 0.273 | 0.136 | -1.247 | |
| ENSMUSG00000069601 | E100 | 6.7339555 | 0.0030706728 | 0.3207887165 | 0.85626216 | 10 | 69830202 | 69830276 | 75 | + | 0.936 | 0.815 | -0.464 |
| ENSMUSG00000069601 | E101 | 1.4807125 | 0.0105199117 | 0.9938791649 | 1.00000000 | 10 | 69830277 | 69831070 | 794 | + | 0.392 | 0.394 | 0.008 |
| ENSMUSG00000069601 | E102 | 10.6883085 | 0.0020263866 | 0.6527423671 | 0.95668705 | 10 | 69833925 | 69834056 | 132 | + | 1.085 | 1.040 | -0.164 |
| ENSMUSG00000069601 | E103 | 11.5576575 | 0.0194290666 | 0.2434023645 | 0.80442163 | 10 | 69835185 | 69835328 | 144 | + | 1.029 | 1.166 | 0.494 |
| ENSMUSG00000069601 | E104 | 0.8216586 | 0.0179286003 | 0.4889450619 | 0.92579672 | 10 | 69835329 | 69835533 | 205 | + | 0.201 | 0.324 | 0.914 |
| ENSMUSG00000069601 | E105 | 13.6753417 | 0.0032098634 | 0.4948438977 | 0.92674629 | 10 | 69837753 | 69838061 | 309 | + | 1.134 | 1.197 | 0.225 |
| ENSMUSG00000069601 | E106 | 8.2168680 | 0.0052507564 | 0.7505853068 | 0.97541370 | 10 | 69838062 | 69838340 | 279 | + | 0.978 | 0.942 | -0.136 |
| ENSMUSG00000069601 | E107 | 7.5426158 | 0.0398907695 | 0.3196169354 | 0.85538296 | 10 | 69838341 | 69838431 | 91 | + | 0.991 | 0.839 | -0.574 |
| ENSMUSG00000069601 | E108 | 38.9807242 | 0.0005207099 | 0.0453345965 | 0.47435072 | 10 | 69840440 | 69840821 | 382 | + | 1.548 | 1.656 | 0.366 |
| ENSMUSG00000069601 | E109 | 0.4992290 | 0.2918223942 | 0.7598450892 | 10 | 69842630 | 69842632 | 3 | + | 0.201 | 0.136 | -0.672 | |
| ENSMUSG00000069601 | E110 | 0.6601124 | 0.0192790232 | 0.8231993140 | 10 | 69842633 | 69842683 | 51 | + | 0.201 | 0.240 | 0.329 | |
| ENSMUSG00000069601 | E111 | 0.0000000 | 10 | 69844608 | 69844694 | 87 | + | ||||||
| ENSMUSG00000069601 | E112 | 0.0000000 | 10 | 69849802 | 69849885 | 84 | + | ||||||
| ENSMUSG00000069601 | E113 | 0.0000000 | 10 | 69850020 | 69850076 | 57 | + | ||||||
| ENSMUSG00000069601 | E114 | 1.8087409 | 0.3064165404 | 0.6382706078 | 0.95668705 | 10 | 69851398 | 69851400 | 3 | + | 0.393 | 0.506 | 0.583 |
| ENSMUSG00000069601 | E115 | 7.8829777 | 0.0024446701 | 0.1751784664 | 0.75672932 | 10 | 69851401 | 69851483 | 83 | + | 0.873 | 1.025 | 0.570 |
| ENSMUSG00000069601 | E116 | 0.1659033 | 0.0345408760 | 0.3947101960 | 10 | 69851484 | 69851635 | 152 | + | 0.000 | 0.136 | 13.043 | |
| ENSMUSG00000069601 | E117 | 95.7340718 | 0.0011622138 | 0.3875018457 | 0.88612996 | 10 | 69859662 | 69863268 | 3607 | + | 1.969 | 1.999 | 0.101 |
Help
Please Click HERE to learn more details about the results from DEXseq.